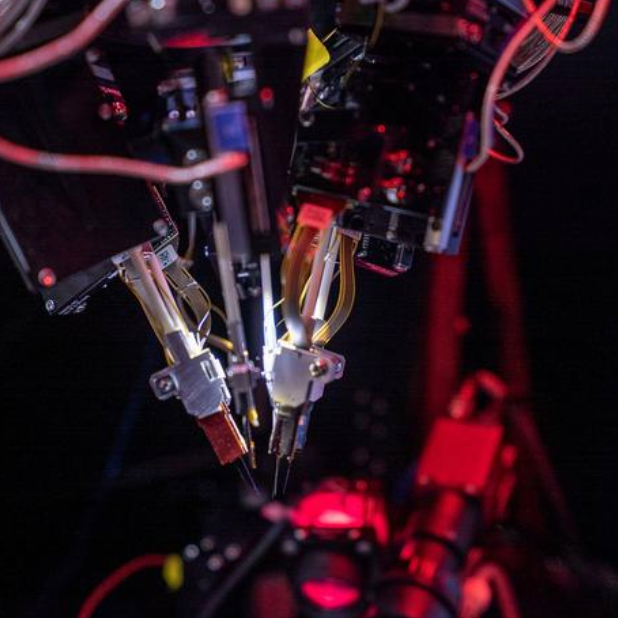
An international team of scientists has found that twisted carbon nanotubes can store high densities of energy to power sensors and other technologies.
The researchers suggest that this finding could advance carbon nanotubes as a promising solution for storing energy in devices that need to be lightweight, compact, and safe; and could also work in a wide range of futuristic technologies, such as space elevators.
Single-walled carbon nanotubes
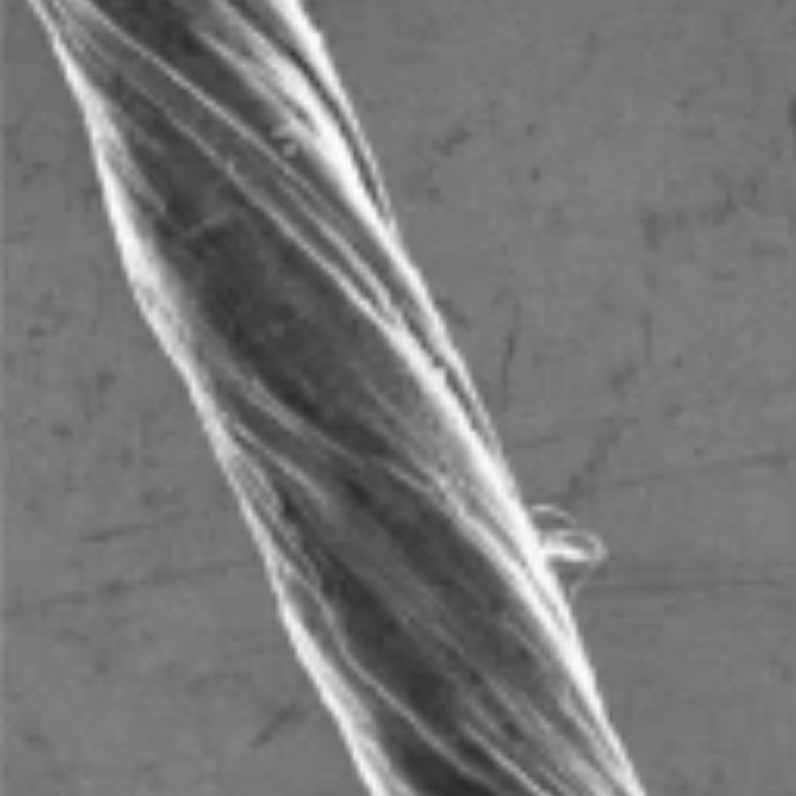
The researchers at the University of Maryland Baltimore County studied single-walled carbon nanotubes. These are like straws, made from pure carbon sheets that are only one-atom thick. They are lightweight, relatively easy to manufacture and about 100 times stronger than steel, as recently described in an open-access Nature Nanotechnology paper.
To investigate carbon nanotubes’ potential for storing energy, the UMBC researchers and colleagues manufactured carbon nanotube “ropes” from bundles of commercially available nanotubes. After pulling and twisting the tubes into a single thread, they coated the ropes with substances that increased the ropes’ strength and flexibility.
Higher energy storage and safer
By twisting them up and measuring the energy that was released as the ropes unwound, they found that the ropes could store 15,000 times more energy per unit mass than steel springs and about three times more energy than lithium-ion batteries.
They also note that materials in the carbon nanotube ropes are safer for the human body than those used in batteries. This stored energy also remains consistent and accessible at temperatures ranging from -60 to +100 °C (-76 to +212 °F).
The team is currently working to incorporate twisted carbon nanotubes as an energy source for a prototype sensor they are developing.
Citation: Utsumi, S., Ujjain, S. K., Takahashi, S., Shimodomae, R., Yamaura, T., Okuda, R., Kobayashi, R., Takahashi, O., Miyazono, S., Kato, N., Aburamoto, K., Hosoi, Y., Ahuja, P., Furuse, A., Kawamata, Y., Otsuka, H., Fujisawa, K., Hayashi, T., Tománek, D., . . . Kaneko, K. (2024). Giant nanomechanical energy storage capacity in twisted single-walled carbon nanotube ropes. Nature Nanotechnology, 1-9. 10.1038/s41565-024-01645-x (open access)
Let us know your thoughts! Sign up for a Mindplex account now, join our Telegram, or follow us on Twitter.

.png)

.png)


.png)